MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

Ribosome Profiling in the Model Diatom Thalassiosira pseudonana - Pichler - 2023 - Current Protocols - Wiley Online Library
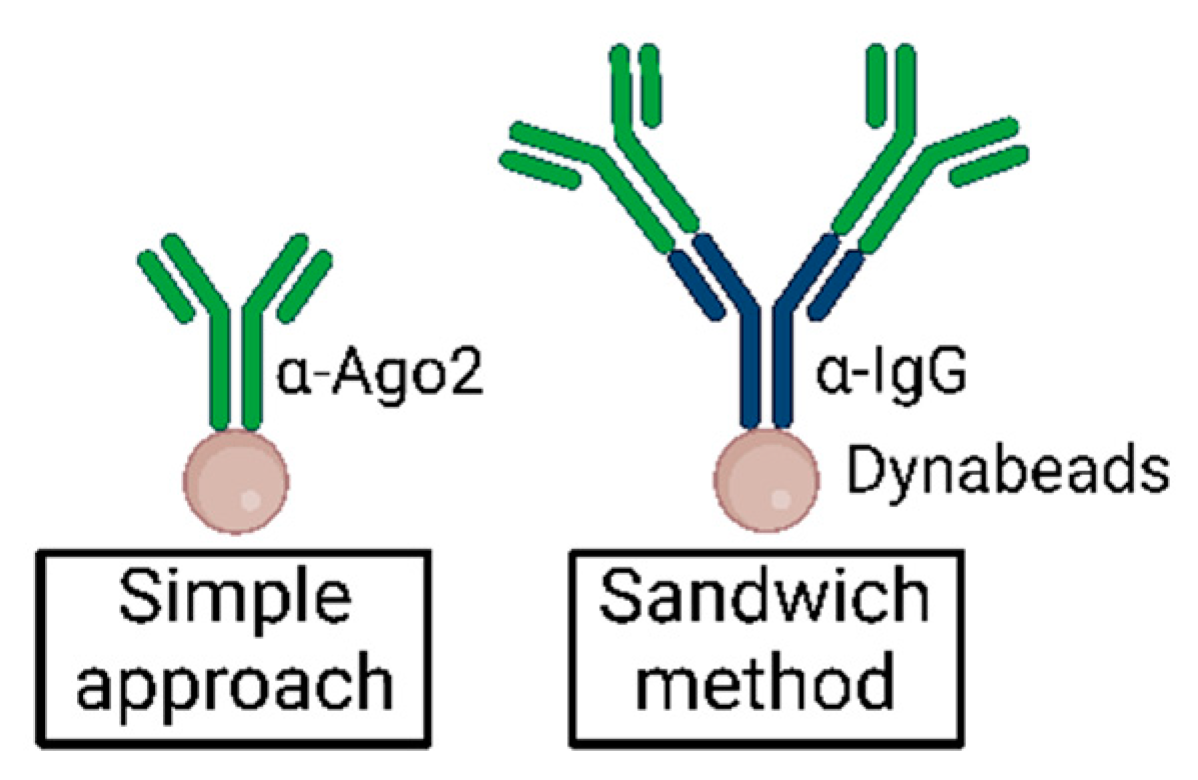
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

Transcriptome-wide identification of RNA-binding protein binding sites using seCLIP-seq | Nature Protocols
HiFi sequencing and software v11.0 release: Technical overview for Sequel II and Sequel IIe system users

Application of a Spacer-nick Gene-targeting Approach to Repair Disease-causing Mutations with Increased Safety

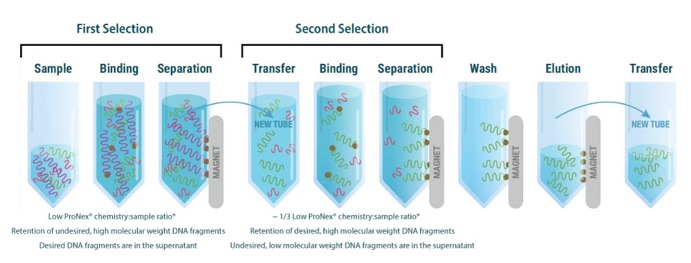
Optimising library size selection for iCLIP2: Sample-to-ProNex bead... | Download Scientific Diagram